Python and its most popular data wrangling library, Pandas, are soaring in popularity. Compared to competitors like Java, Python and Pandas make data exploration and transformation simple.
But both Python and Pandas are known to have issues around scalability and efficiency.
Python loses some efficiency right off the bat because it’s an interpreted, dynamically typed language. But more importantly, Python has always focused on simplicity and readability over raw power. Similarly, Pandas focuses on offering a simple, high-level API, largely ignoring performance. In fact, the creator of Pandas wrote “The 10 things I hate about pandas,” which summarizes these issues:

So it’s no surprise that many developers are trying to add more power to Python and Pandas in various ways. Some of the most notable projects are:
- Dask: a low-level scheduler and a high-level partial Pandas replacement, geared toward running code on compute clusters.
- Ray: a low-level framework for parallelizing Python code across processors or clusters.
- Modin: a drop-in replacement for Pandas, powered by either Dask or Ray.
- Vaex: a partial Pandas replacement that uses lazy evaluation and memory mapping to allow developers to work with large datasets on standard machines.
- RAPIDS: a collection of data-science libraries that run on GPUs and include cuDF, a partial replacement for Pandas.
There are others, too. Below is an overview of the Python data wrangling landscape:
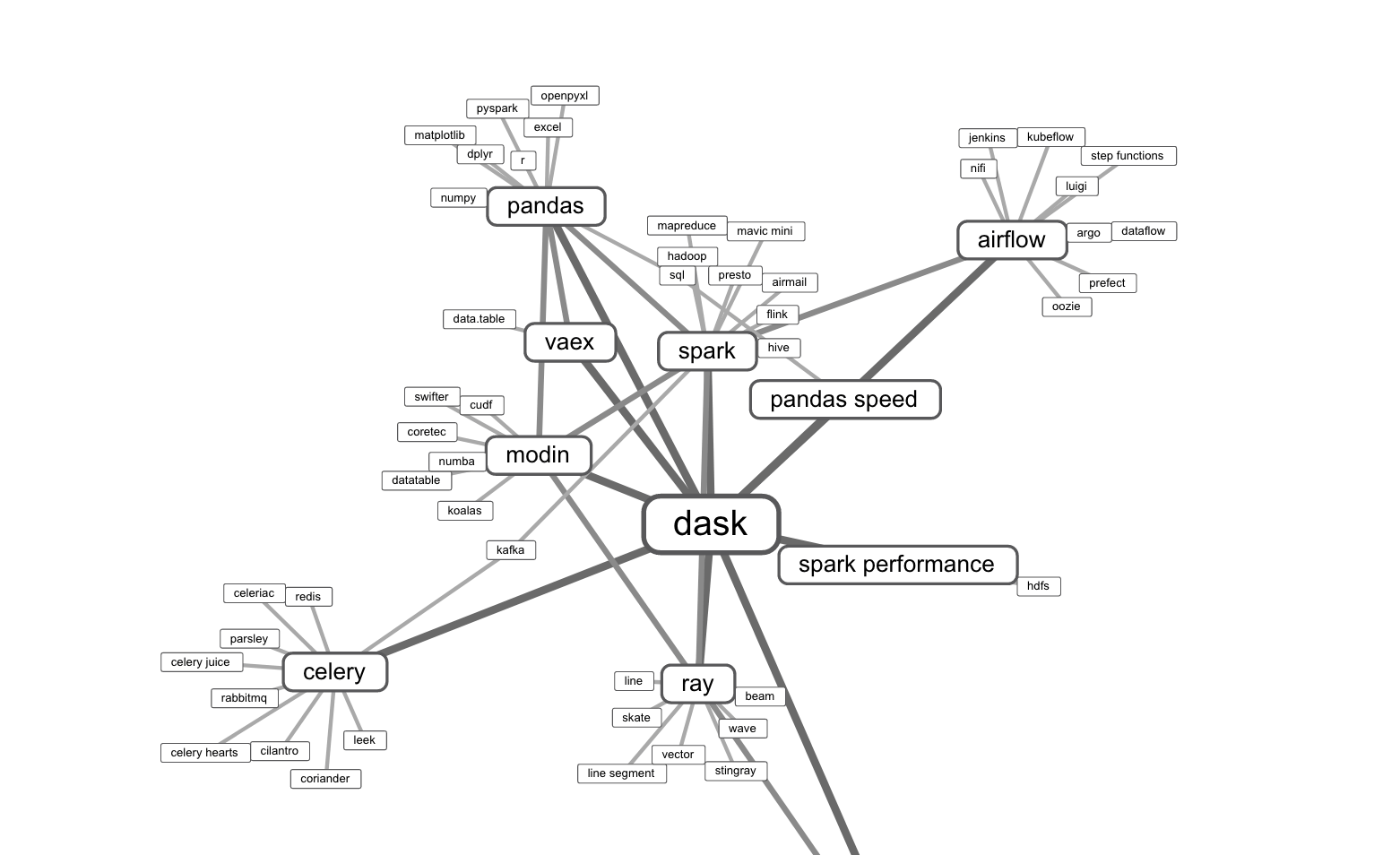
So if you’re working with a lot of data and need faster results, which should you use?
Just tell me which one to try
Before you can make a decision about which tool to use, it’s good to have some more context about each of their approaches. We’ll compare each of them closely, but you’ll probably want to try them out in the following order:
- Modin, with Ray as a backend. By installing these, you might see significant benefit by changing just a single line (`import pandas as pd` to `import modin.pandas as pd`). Unlike the other tools, Modin aims to reach full compatibility with Pandas.
- Dask, a larger and hence more complicated project. But Dask also provides Dask.dataframe, a higher-level, Pandas-like library that can help you deal with out-of-core datasets.
- Vaex, which is designed to help you work with large data on a standard laptop. Its Pandas replacement covers some of the Pandas API, but it’s more focused on exploration and visualization.
- RAPIDS, if you have access to NVIDIA graphics cards.
Quick comparison
Each of the libraries we examine has different strengths, weaknesses, and scaling strategies. The following table gives a broad overview of these. Of course, as with many things, most of the scores below are heavily dependent on your exact situation.

These are subjective grades, and they may vary widely given your specific circumstances. When assigning these grades, we considered:
- Maturity: The time since the first commit and the number of commits.
- Popularity: The number of GitHub stars.
- Ease of Adoption: The amount of knowledge expected from users, presumed hardware resources, and ease of installation.
- Scaling ability: The broad dataset size limits for each tool, depending on whether it relies mainly on RAM, hard drive space on a single machine, or can scale up to clusters of machines.
- Use case: Whether the libraries are designed to speed up Python software in general (“General”), are focused on data science and machine learning (“Data science”), or are limited to simply replacing Pandas’ ‘DataFrame’ functionality (“DataFrame”).
CPUs, GPUs, Clusters, or Algorithms?
If your dataset is too large to work with efficiently on a single machine, your main options are to run your code across…
- ...multiple threads or processors: Modern CPUs have several independent cores, and each core can run many threads. Ensuring that your program uses all the potential processing power by parallelizing across cores is often the easiest place to start.
- ...GPU cores: Graphics cards were originally designed to efficiently carry out basic operations on millions of pixels in parallel. However, developers soon saw other uses for this power, and “GP-GPU” (general processing on a graphics processing unit) is now a popular way to speed up code that relies heavily on matrix manipulations.
- ...compute clusters: Once you hit the limits of a single machine, you need a networked cluster of machines, working cooperatively.
Apart from adding more hardware resources, clever algorithms can also improve efficiency. Tools like Vaex rely heavily on lazy evaluation (not doing any computation until it’s certain the results are needed) and memory mapping (treating files on hard drives as if they were loaded into RAM).
None of these strategies is inherently better than the others, and you should choose the one that suits your specific problem.
Parallel programming (no matter whether you’re using threads, CPU cores, GPUs, or clusters) offers many benefits, but it’s also quite complex, and it makes tasks such as debugging far more difficult.
Modern libraries can hide some – but not all – of this added complexity. No matter which tools you use, you’ll run the risk of expecting everything to work out neatly (below left), but getting chaos instead (below right).

Dask vs. Ray vs. Modin vs. Vaex vs. RAPIDS
While not all of these libraries are direct alternatives to each other, it’s useful to compare them each head-to-head when deciding which one(s) to use for a project.
Before getting into the details, note that:
- RAPIDS is a collection of libraries. For this comparison, we consider only the cuDF component, which is the RAPIDS equivalent of Pandas.
- Dask is better thought of as two projects: a low-level Python scheduler (similar in some ways to Ray) and a higher-level Dataframe module (similar in many ways to Pandas).
Dask vs. Ray
Dask (as a lower-level scheduler) and Ray overlap quite a bit in their goal of making it easier to execute Python code in parallel across clusters of machines. Dask focuses more on the data science world, providing higher-level APIs that in turn provide partial replacements for Pandas, NumPy, and scikit-learn, in addition to a low-level scheduling and cluster management framework.
The creators of Dask and Ray discuss how the libraries compare in this GitHub thread, and they conclude that the scheduling strategy is one of the key differentiators. Dask uses a centralized scheduler to share work across multiple cores, while Ray uses distributed bottom-up scheduling.
Dask vs. Modin
Dask (the higher-level Dataframe) acknowledges the limitations of the Pandas API, and while it partially emulates this for familiarity, it doesn’t aim for full Pandas compatibility. If you have complicated existing Pandas code, it’s unlikely that you can simply switch out Pandas for Dask.Dataframe and have everything work as expected. By contrast, this is exactly the goal Modin is working toward: 100% coverage of Pandas. Modin can run on top of Dask but was originally built to work with Ray, and that integration remains more mature.
Dask vs. Vaex
Dask (Dataframe) is not fully compatible with Pandas, but it’s pretty close. These close ties mean that Dask also carries some of the baggage inherent to Pandas. Vaex deviates more from Pandas (although for basic operations, like reading data and computing summary statistics, it’s very similar) and therefore is also less constrained by it.
Ultimately, Dask is more focused on letting you scale your code to compute clusters, while Vaex makes it easier to work with large datasets on a single machine. Vaex also provides features to help you easily visualize and plot large datasets, while Dask focuses more on data processing and wrangling.
Dask vs. RAPIDS (cuDF)
Dask and RAPIDS play nicely together via an integration provided by RAPIDS. If you have a compute cluster, you should use Dask. If you have an NVIDIA graphics card, you should use RAPIDS. If you have a compute cluster of NVIDIA GPUs, you should use both.
Ray vs. Modin or Vaex or RAPIDS
It’s not that meaningful to compare Ray to Modin, Vaex, or RAPIDS. Unlike the other libraries, Ray doesn’t offer high-level APIs or a Pandas equivalent. Instead, Ray powers Modin and integrates with RAPIDS in a similar way to Dask.
Modin vs. Vaex
As with the Dask and Vaex comparison, Modin’s goal is to provide a full Pandas replacement, while Vaex deviates more from Pandas. Modin should be your first port of call if you’re looking for a quick way to speed up existing Pandas code, while Vaex is more likely to be interesting for new projects or specific use cases (especially visualizing large datasets on a single machine).
Modin vs. RAPIDS (cuDF)
Modin scales Pandas code by using many CPU cores, via Ray or Dask. RAPIDS scales Pandas code by running it on GPUs. If you have GPUs available, give RAPIDS a try. But the easiest win is likely to come from Modin, and you should probably turn to RAPIDS only after you’ve tried Modin first.
Vaex vs. RAPIDS (cuDF)
Vaex and RAPIDS are similar in that they can both provide performance boosts on a single machine: Vaex by better utilizing your computer’s hard drive and processor cores, and RAPIDS by using your computer’s GPU (if it’s available and compatible). The RAPIDS project as a whole aims to be much broader than Vaex, letting you do machine learning end-to-end without the data leaving your GPU. Vaex is better for prototyping and data exploration, letting you explore large datasets on consumer-grade machines.
Final remarks: Premature optimization is the root of all evil
It’s fun to play with new, specialized tools. That said, many projects suffer from over-engineering and premature optimization. If you haven’t run into scaling or efficiency problems yet, there’s nothing wrong with using Python and Pandas on their own. They are widely used and offer maturity and stability, along with simplicity.
You should only start looking into the libraries discussed here once you’ve reached the limitations of Python and Pandas on their own. Otherwise you risk spending too much time choosing and configuring libraries instead of making progress on your project.
At DataRevenue, we’ve built many projects with these libraries and know when and how to use them. If you need a second opinion, reach out to us. We’re happy to help.