Metabolomics is a new field. As more accurate instruments have become available, the field has grown steadily over the last 20 years. One metabolomics analysis platform in particular, liquid chromatography coupled mass spectrometry (LCMS), has seen massive development. It is quickly becoming the driving force of clinical diagnostics.
Every year, researchers publish hundreds of papers about using metabolomics to address human health. However, clinical applications are noticeably lacking in comparison with this publication rate. This suggests there are significant challenges in using metabolomics for diagnostics.
We interviewed Oliver Fiehn, director of the UC Davis West Coast Metabolomics Center, a world-leading institute for metabolomics analysis. Oliver has published over 340 articles on the use, development, and outlook of metabolomics for human health. In this conversation we learned how many new tools and techniques still fall short of clinical goals, and how to best prepare for the “valley of death” from benchtop to clinic.
Metabolomics creates new viewpoints for big data
Metabolomics uses large sample sizes and high-quality comparisons to streamline diagnosis and discover biomarkers from complex samples. “There's an endless possibility for applications of metabolomics,” Oliver says. And there is no single way to look at the data. Many “omics” fields rely on multiple analyses in conjunction to arrive at a question’s true answer. “If you [take] say, iceberg-omics, you could take a photograph of the iceberg above sea level and say, ‘I have an iceberg and it's that big.’ But if you have another tool that can also look below the sea level, you can see it's much bigger. Metabolomics is not just about a type of instrument, or one protocol. It's the combination of everything in the workflow that makes it standardized and reliable.”
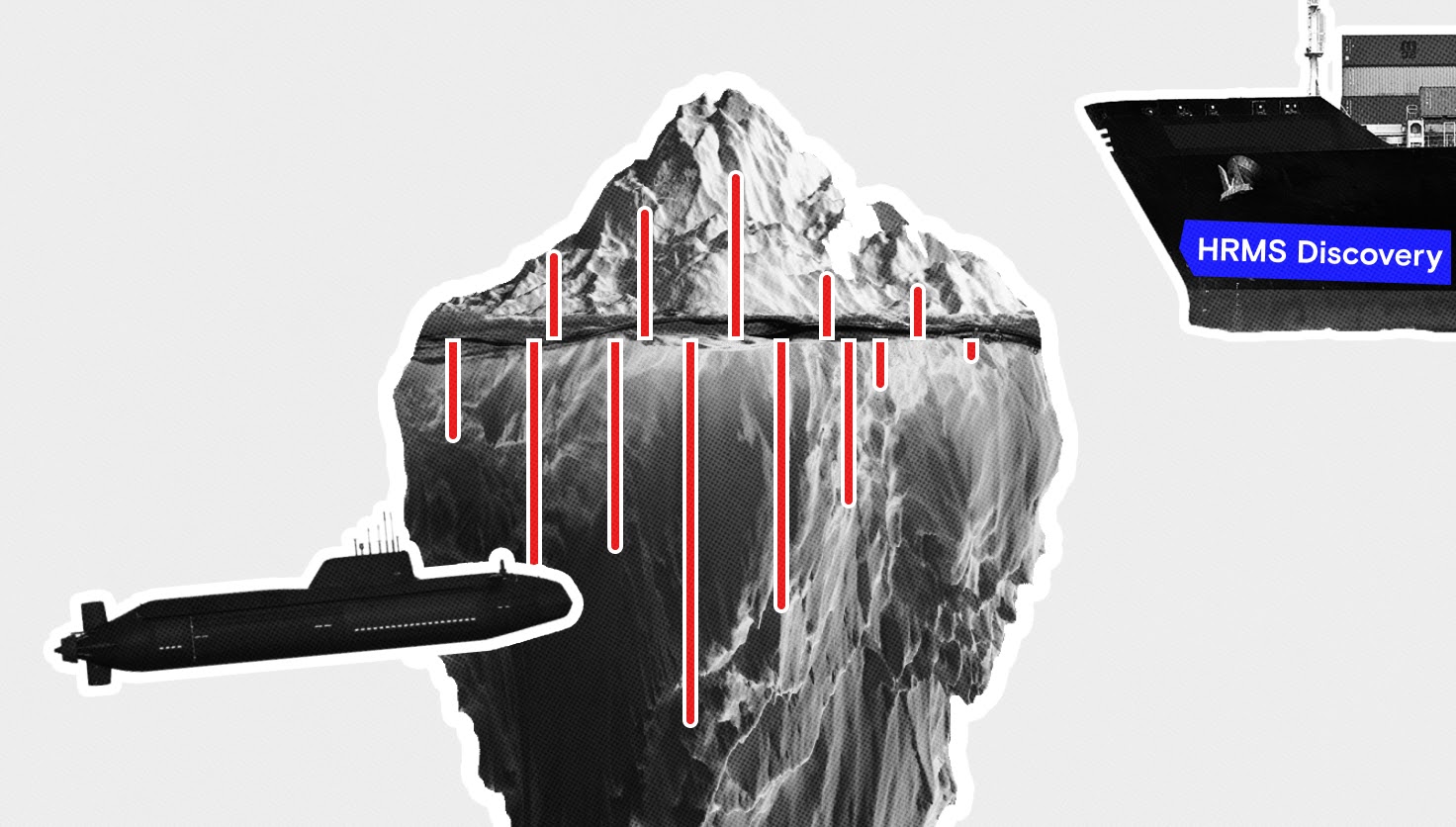

Innovations in metabolomics include ways to improve sample throughput, new software and algorithm development, new experimental workflows, and new techniques for biomarker discovery. These could all translate to better patient care in the clinic.
Publication to patient: It’s a long journey
Many different LCMS metabolomics workflows have been developed by academics over the last 20 years. But overall, clinics have been slow to adopt metabolomic techniques. From Oliver’s perspective, this is because academia and clinical practice are two separate things, each motivated by different objectives. “One thing is academia, where we publish a paper and we are very proud of what the paper says. It is another thing in the clinics where an actual doctor has an actual patient. That's very different.”
Many untargeted approaches focus on a discovery model – i.e., what is new? – rather than a diagnostics one – i.e., does this imply disease? This creates a disconnect between algorithm development and clinical use cases that can’t always be fixed.
Although every researcher hopes innovations at the bench will connect directly to the clinic, there are great challenges. Or as Oliver puts it, “there's a valley of death in between” the two.
Oliver makes a comparison with drug discovery, in which the discovery-to-drug process regularly takes over a decade: “People underestimate the trouble it takes to get new therapies not only approved, but accepted in the field.” Drug development has a high rate of attrition: Over 98% of drug candidates will fail clinical trials.
Based on clinical adoption, we can expect a comparable attrition rate for new metabolomics analyses. What kinds of challenges should a researcher expect to overcome for their innovations to reach the clinic?
Metabolomics tools should be more adaptable
Broad applicability is frequently a secondary design objective for academic metabolomics pipelines, if it’s considered at all. Pipelines are often custom designed to a single lab’s workflow.
Academic research is often at the forefront of tool creation, so there are implicit challenges for widespread use:
- Processes are often code-based: Researchers write customized code, requiring all its users to understand how to read and use it.
- Data formats are a pain point: Pipelines often require specific data formatting. Some of this is proprietary and can’t be easily converted.
- Study design and patient diagnosis are not the same thing: Untargeted metabolomics can lead to new discoveries. But clinical uses focus on targeted detection in complex biological samples.
Oliver hints that even the best studies have a big gap between study design and real-world application. Research software is usually tailored toward discovery, for example, of new molecules. The viewpoints these tools offer can be very helpful to an academic looking for new molecules. But they’re of little use to a clinician: “That's not how [clinicians] look at things. They say, ‘Give me a tool that enables making a decision, or a diagnostic.’”
That doesn’t mean these tools won’t be useful in the clinic. But it means there’s a need to frame them for the problems clinicians face.
Building clinical relevance is the key to adoption
In a clinical setting the questions become, “What is wrong with this person? How do I help? How can I improve my therapies? How do I make a multi-therapy option?” To evaluate whether tools and workflows are suitable for clinical adoption, it helps to look at why they were developed in the first place. Do they make a new diagnosis possible, or speed up sample processing time?
As Oliver puts it, “Nothing is as successful as a success. I always like case studies where they show that something worked because of a piece of software that was deployed, and it likely could have not been done by any other software.”
Robust case studies show that new methodologies can handle complex questions in real systems. And as more labs adopt a methodology because of its ease of use and robust results, this bolsters trust, whether for a new program or experimental setup.
If a new utility can handle a complex question in a real system while minimizing the pain points of learning a new software or coding language, the chances for its adoption and widespread acceptance are much greater. The UC Davis Metabolomics Center, the Metabolomics Association of North America, and the UC San Diego Center for Computational Mass Spectrometry all offer workshops and training sessions to help alleviate the stress of adopting new methodologies.
Low cost and fast analyses
When designing a clinical assessment, material cost and time are huge factors. To develop an approach that is practical on scale, Oliver asks “What can I do in five minutes?"
In clinical diagnostics, the focus is on sample throughput and monitoring patient response. And while metabolomics could revolutionize the landscape of modern medicine, new platforms must first cross the “valley of death” from lab-bench to patient. For this to happen, analyses need to be quick, cheap to perform, reproducible across many labs, and practical for use on human samples.
“If I have 200,000 samples, you can't do a one hour assay – that's not going to be cost effective.” Worse, some treatments may be too late if an assay takes hours. In the real world, time can cost lives.
Working with the Department of Defense, Oliver and his team are developing a rapid assay to determine a crucial treatment for soldiers on the front line: Head trauma and burn injuries often co-occur but have opposing treatments, and choosing the wrong one can be fatal. As Oliver explains, “if you have a severe burn, you may need to drink a lot of glucose+electrolyte water – but if you have head trauma, such treatment may lead to brain swelling.” An assay that can assess a patient's profile in five minutes and help make the right choice could make a life-saving difference.
Treating every patient as a unique sample set
Metabolomics studies can be rigidly structured, with careful controls and large sample sizes. This is an unrealistic expectation in a clinical setting. As Oliver puts it, “the problem in clinical metabolomics is that you have individual patients. You don't have a cohort of 1,000 people coming into the clinic to analyze.”
So can we translate a single patient’s profile into real knowledge about what might be going on?
Creating individual patient profiles by collecting longitudinal data could hold new opportunities. Instead of comparing large cohorts, patients could be studied over longer periods of time, optimally starting before any diseases develop. During a disease treatment, you would “ping the system” of the patient through a treatment and compare the response over time. With enough of these longitudinal datasets, valuable information can be obtained and applied to diagnosis and treatment in the clinic.
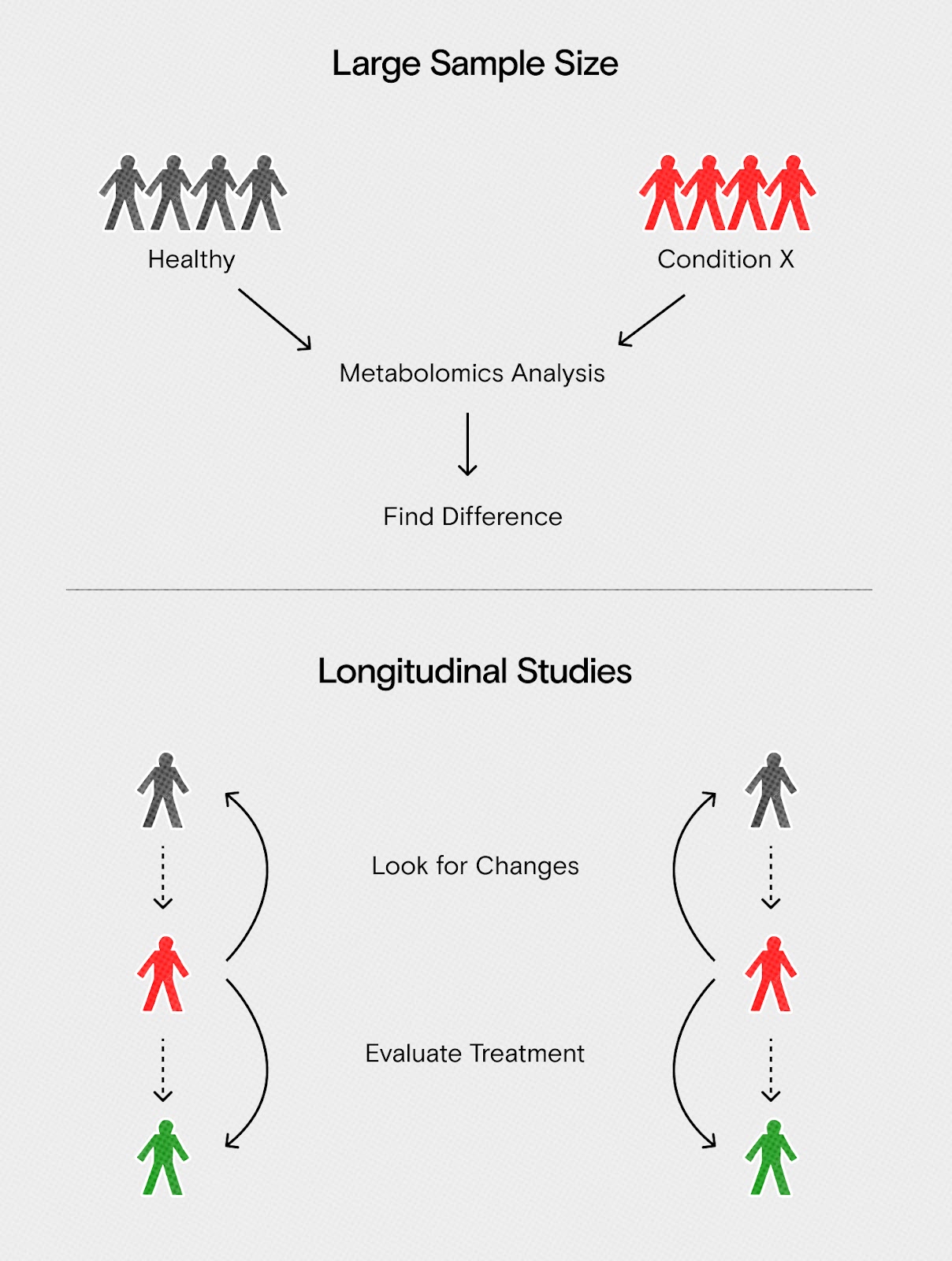
The NIH recently launched the “All of Us” initiative to collect longitudinal data on one million Americans, including healthy individuals. These data are a valuable link between the lab bench and the clinic. As health conditions appear, scientists can look at a patient’s profile for new disease biomarkers, evaluate treatment effectiveness, or study long-term changes.
“The NIH will put out these huge cohorts so that we have comparative datasets. That way, people who use AI/ML methods would be able to make sense out of the data,” says Oliver. This could enable new and rapid diagnosis technologies. “Then, if the person who's sick walks into the door in the clinic in Sacramento, Oliver Fiehn with his tool could detect” and diagnose the disease.
Metabolomic studies can focus on a variety of clinical data. If researchers think about these data in new ways, they'll discover new ways to transition to the clinic. There's an endless possibility of applications, but we still need to keep the realistic use cases in mind.
Learn more about taking your research from bench to clinic
- Check out the UC Davis Metabolomics Center
- Sign up for their metabolomics workshops
- Learn more about solving challenges with translational metabolomics
- Discover more about the NIH All of Us initiative
We have experience in translating metabolomics data analysis for clinical use. Schedule a consultation with us to learn how to scale up your metabolomics workflow, or optimize it for clinical use.